

Nucleotide excision repair. Atlas of Genetics and Cytogenetics in Oncology and Haematology. Home Genes. Solid Tumours. Cancer- Prone. Deep Insight. Case Reports. XY1.
NA. Leon H. F. Mullenders (1), Anne Stary (2) and Alain Sarasin (2). Department. of Radiation Genetics and Chemical Mutagenesis, - MGC Leiden. University Medical Center, P. O. Box 9. 50. 3, 2.
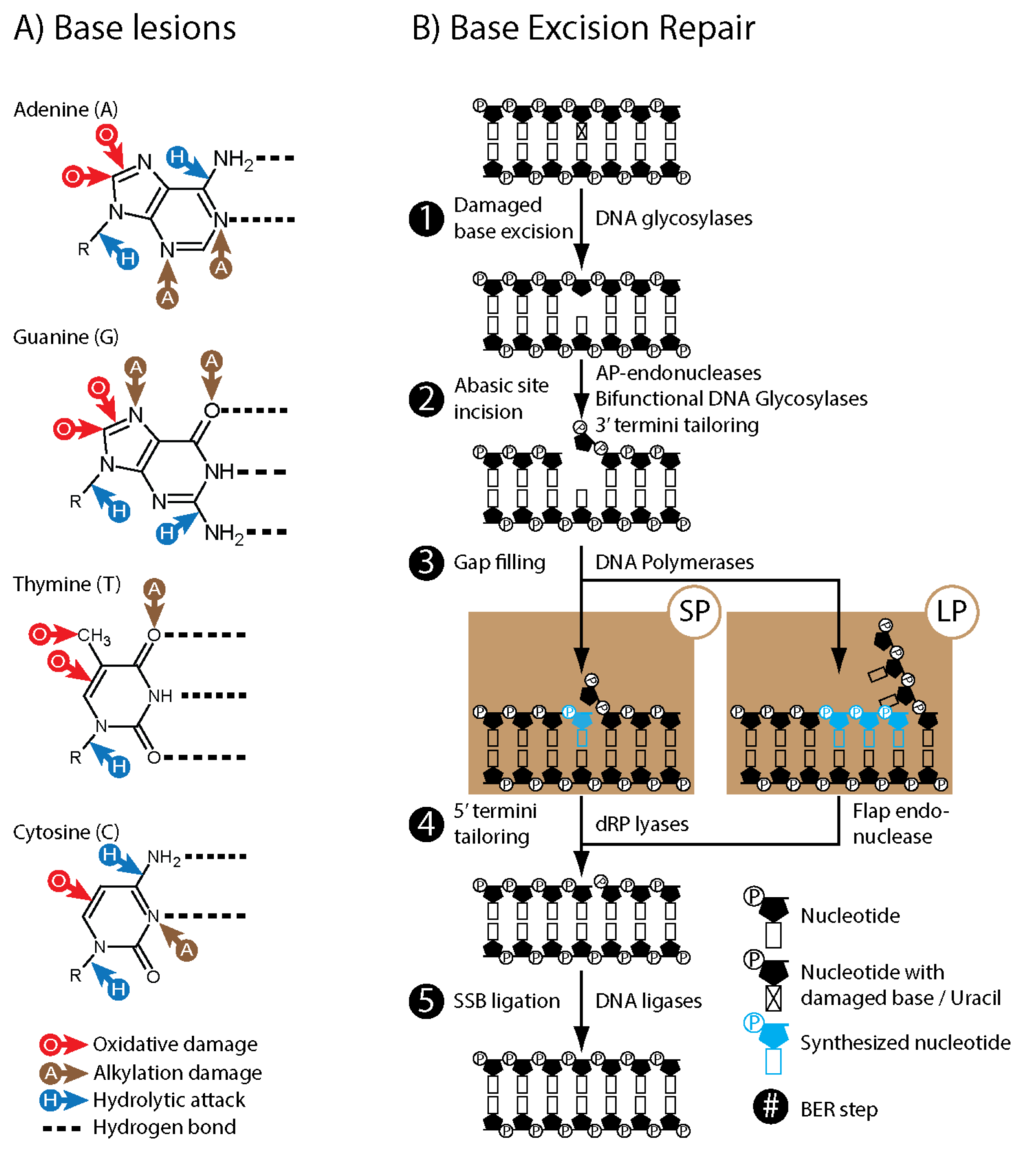
- Base excision repair of DNA in mammalian cells.
- The second defense line is to fix the oxidative damage through one of the DNA repair pathways (e.g.
- Base excision repair: the long and short of it. Base excision repair, glycosylase, DNA damage, alkylation. The one-nucleotide short-patch BER pathway is completed by the action of DNA ligase III.
- DNA repair in mammalian cells: Base excision repair. The shortpatch BER pathway leads to a repair tract of a single nucleotide.
RA Leiden and J. A. Cohen. Institute, Interuniversity Research Institute for Radiopathology and.
Long Patch Base Excision Repair with Purified Human Proteins. Base excision repair (BER) pathway, protects both nuclear and mitochondrial DNA from 'spontaneous DNA damage', mainly generated by eactive oxigen spices (ROS) produced by the normal metabolism of the cell.
Radiation Protection, Leiden, The Netherlands. Laboratory of Genetic Instability and Cancer, UPR 2. CNRS, BP 8. 9. 48.
Villejuif, France. February 2. 00. 1. All living organisms are equipped with DNA repair systems that can cope. DNA lesions. NER is a complex process in which basically the. DNA lesion. (ii) separation of the double helix at the DNA lesion site.
DNA fragment. (v) DNA repair synthesis to replace the gap and. Patients characteristically show severe photosensitivity and. Bootsma et al., 1. Complementation studies revealed that eight genes are. XP: XPA through XPG and XPV (XP- Variant). Mutations in the XP.
XP- variant) lead to defective NER and hypersensitivity to UV. Normally, the. translesion synthesis is carried out by the polymerase eta, which is. XP variant. XP- V patients are more UV- sensitive than normal.
XP. They develop skin cancers around. Cockayne' syndrome (CS) is a rare disorder that is associated with a wide.
Beside other symptoms, the patients generally. In contrast to XP. CS is not associated with an enhanced incidence of skin cancer. Cells from. CS patients are hypersensitive to the cytotoxic effects of UV and are.
UV inhibited DNA and RNA. Two CS complementation groups (A and B) have been established. A. third group encompasses patients exhibiting both XP and CS symptoms, they. XP groups B, D or G. The progressive neurological abnormalities. CS may be due to the inability of CS cells to repair. DNA lesions (Le. Page et al., 2.
The first. pathway is global genome repair (GGR) and involves repair activity that. DNA lesions across the genome. Although the efficiency of this.
A second NER pathway is coupled. This. pathway involves repair activity that is directed to the transcribed strand. Two proteins have been identified and. NER, i. e. In addition, the XPE protein has been shown to have a. DNA, but whether it is required for the damage. NER remains unclear. Cells from XPA patients are.
UV and have very low nucleotide excision repair. In vitro the XPA protein binds preferentially to damaged DNA.
DNA. The XPA protein binds to replication protein A. RPA) which enhances the affinity of XPA for damaged DNA and is essential. NER. The other complex that has been implicated in DNA damage. XPC- HR2. 3B. XPC cells have low NER repair capacity, but the. Binding of XPC- HR2. B to a DNA lesion.
XPA protein can bind and the whole. This would imply that. XPA protein has binding affinity for other repair proteins. Indeed, the. XPA protein has been shown to bind to ERCC1 and TFIIH. The XPC- HR2. 3B. In case of transcription. RNA polymerase is stalled at a lesion, the DNA is.
XPA can bind independently of XPC- . HR2. 3B complex. The function of the gene. Band shift assays suggested that.
XPE gene product acts as a damaged DNA binding protein (DDB), with high. UV- induced 6- 4. PP. However, defective DDB binding activity is. XPE mutant cell lines and in fact two (or even. In. cells from several XPE patient mutations in p. XPE cells are not.
CPD accessible to the NER machinery, but is not. PP. Interestingly, cell lines and primary tissues. Tang. et al., 2. 00.
This explains the absence of GGR of CPD in these cells. The striking discovery that subunits of basal. TFIIH were involved in NER sheds light on a new aspect. NER : a close coupling to transcription via common use of essential. Two repair proteins, encoded by XPB and XPD genes, appeared to be. TFIIH, a large. complex involved in the initiation of transcription. The XPB and XPD. proteins displayed 3'- 5' and 5'- 3' helicase activity respectively.
Schaeffer et al., 1. TFIIH fulfills a dual role in transcription. NER and the role of TFIIH in NER might closely mimic its. In transcription initiation.
TFIIH is thought to be involved in unwinding of the promoter site and to. In the NER process TFIIH causes unwinding of the. XPC- HR2. 3B and XPA- RPA. NER proteins around the damaged site. The helicase activity of XPD is. NER but not for transcription initiation. So , there is. much more XPD patients, and only two patients have been described as XP and.
CS. The XPF protein and the ERCC1 protein form a complex that. NER reaction. XPF- ERCC1 also binds to XPA. ERCC1) and to RPA (through XPF) but not preferentially to damaged. DNA. The XPG protein has DNA endonuclease activity without preference for. DNA and is responsible for the 3' incision made during NER.
At the. site of a lesion NER proteins create a DNA bubble structure over a length. XPG protein incises the damaged DNA. DNA- ds. DNA junction.
In most studies the. XPG protein appeared to be made prior to and. XPF- ERCC1. Patients belonging to the. XP- G complementation group clinically exhibit heterogeneous symptoms, from.
CS. XP- G cells are almost. UV- sensitive as XP- A cells. About half. of the described XPG patients exhibit also CS symptoms. In contrast to XPG. XP- F patients have a relatively mild XP phenotype without neurological.
Cells from XP- F patients are slightly UV- sensitive and. UV- irradiation. Proliferating Cell Nuclear.
Antigen (PCNA) is required for DNA synthesis by DNA polymerases delta and. PCNA has also been shown to be required for NER in vitro i. Biochemical analysis and fluorescence microscopy. UV- irradiation PCNA (that usually. The enzymes. involved in these pathways are normal in DNA repair- deficient cells. It is conceivable that the damage recognition.
DNA lesions will lead to more rapid repair. The lesion. recognition and binding potency of proteins that are involved in damage. DNA lesion itself or.
DNA helical structure. Some lesions such as. PP and CPD, are large bulky lesions located in. DNA helix and are recognized by NER proteins as.
DNA. DNA is thought to be a dynamic. Lesions that interfere with these. DNA may be recognized by repair proteins. Lesions. that have been shown to be a good substrate for NER often cause local.
DNA bases around the damaged site. UV- induced CPD as. NER when they are superimposed on a mismatch than in normally. DNA. The unwinding of a few basepairs energetically favours.
DNA and this may facilitate further unwinding by NER. Repair of DNA lesions that are substrates for NER by themselves.
The latter are known to consist of. DNA sequences and their chromatin structure is. DNAse. 1. Thus, the. DNA lesions to. repair proteins. Indeed, when repair was investigated at the nucleotide. Subsequently, TCR was shown to operate in a variety of organisms.
All data indicate that TCR is directly. All. classical XP cells are deficient in TCR except the group C that is fully. GGR but proficient in TCR (Van Hoffen et al., 1.
However. until now it is not clear how repair is coupled to transcription. A major. obstacle that prevents a major breakthrough, is the lack of a cell free.
TCR. In mammalian cells such. Cockayne' syndrome might fulfill such a function. Similarly. to the mdf bacteria strain, Cockayne syndrome cells are unable to perform. The defect in transcription- coupled repair has been. CS cells to restore UV- inhibited RNA synthesis. Mayne and Lehmann 1.
Slow removal of DNA lesions from transcription. Moreover, by analogy to bacteria. NER proteins. Indeed, several investigators. CSB can be copurified with RNA polymerase II but could not. CSB with any other tested NER component. In cells. that have been treated with UV, a small fraction of RNA polymerase II. Bregman et al., 1.
However, neither in CS- A. CS- B cells this specific response was observed. One explanation. favoured by several studies, is that the polymerase could be ubiquitinated. In this model, CS proteins would be required.
 RSS Feed
RSS Feed
